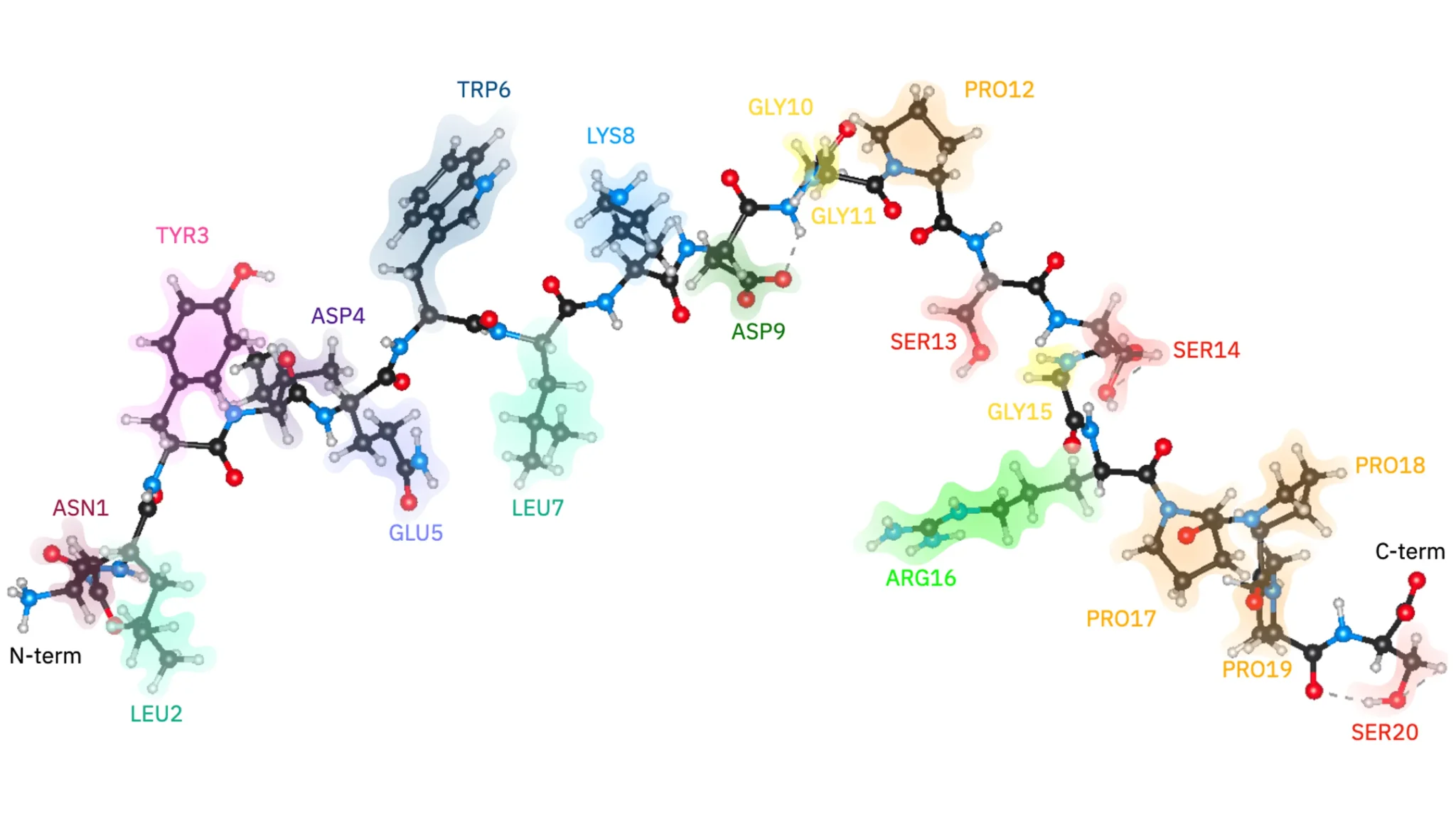
Cleveland Clinic and IBM researchers have successfully simulated the electronic structure of a 303-atom miniprotein, Trp-cage, using a quantum-centric supercomputing workflow and an IBM Quantum Heron r2 system. This achievement addresses a critical limitation in classical computing, where accurately modeling the electronic structure of increasingly large molecules becomes exceptionally challenging; current methods struggle with the complexity of entire proteins. The team fragmented Trp-cage into computationally manageable “clusters” using a technique called wave function-based embedding, then used quantum computing to simulate the most complex of these. “I’m sort of pinching myself that we were able to do it,” said Dr. Kenneth Merz, PhD, suggesting a potential step toward impactful applications in chemical, materials science, and medical research.
Trp-cage Miniprotein Enables First Quantum Electronic Structure Simulation
A joint Cleveland Clinic and IBM research team has achieved the first quantum simulation of a protein’s electronic structure, a feat previously limited by the computational power of classical computers. This advance addresses a fundamental challenge in molecular modeling; the accuracy of electronic structure calculations becomes increasingly challenging as molecule size increases, rendering comprehensive simulations of complex proteins impractical for conventional systems. Mario Motta, a co-author of the study, said that proving this approach works for Trp-cage is a step toward larger molecules, highlighting the scalability potential of the method. The workflow leverages the strengths of both quantum and high-performance classical computing, a core principle of QCSC, where quantum processors contribute to cluster calculations while classical computers handle simpler regions and assemble the complete molecular picture.
This breakthrough builds upon the development of sample-based quantum diagonalization (SQD), an algorithm that efficiently navigates the exponentially growing number of electron configurations in larger molecules. The researchers initially aimed to simulate only a few amino acids, but the workflow’s performance allowed them to scale up to the full Trp-cage structure and obtain meaningful results, potentially leading to future applications in pharmaceutical research and materials science.
Wave Function-Based Embedding Fragments Molecules for Quantum Compute
The pursuit of accurately simulating molecular behavior has long been constrained by the limitations of classical computing, particularly as molecular size and complexity increase; however, a collaborative effort between the Cleveland Clinic and IBM has yielded a quantum-centric supercomputing (QCSC) workflow that offers a potential pathway forward. Researchers successfully modeled the electronic structure of the 303-atom miniprotein Trp-cage, a feat previously unattainable due to the increasing challenge of accurate electronic structure calculations as system size increases. The team fragmented Trp-cage into computationally manageable “clusters” using a technique called wave function-based embedding, then used quantum computing to simulate the clusters. Mario Motta, co-author of the paper, said that the plan was initially to simulate just a couple of amino acids, highlighting the surprising scalability of the workflow.
The resulting data from each cluster is then reassembled to create a complete picture of the molecule’s electronic structure, revealing crucial information about electron distribution and interactions. This approach, where quantum computers work with classical computers in hybrid workflows, is an early look at quantum-centric supercomputing in action. The team’s success builds upon an algorithm called sample-based quantum diagonalization (SQD), which efficiently addresses the challenge of exponentially growing configurations in electronic structure calculations.
We sort of dropped everything. I met with a few people in my group over the weekend, and we decided to just go all in on SQD.
Sample-Based Quantum Diagonalization (SQD) Algorithm Drives QCSC Workflows
Cleveland Clinic and IBM researchers are developing a new approach to molecular simulation, leveraging a hybrid quantum-classical workflow centered around the sample-based quantum diagonalization (SQD) algorithm. This method addresses a fundamental limitation in computational chemistry: the exponential increase in complexity when modeling the electronic structure of larger molecules using traditional computers. While classical methods efficiently handle certain aspects of protein behavior, accurately simulating entire proteins quantum-mechanically has remained impractical until now. The team’s recent success in modeling the 303-atom miniprotein Trp-cage represents a significant step toward overcoming these hurdles and unlocking applications across diverse fields. The workflow relies on wave function-based embedding (EWF) to dissect Trp-cage into manageable “clusters,” each representing a local region around an atom and its entanglement with neighbors.
Crucially, the complexity varies between clusters; those at the protein’s core, enmeshed in complex interactions, are suitable for quantum computation, while simpler, peripheral clusters can be efficiently modeled using classical methods. This division of labor is central to quantum-centric supercomputing (QCSC), where classical and quantum resources work together to solve problems, using the strengths of both paradigms. The quantum computer samples the vast space of possible electron configurations, identifying key areas for the classical computer to refine, ultimately leading to a complete solution for the molecule’s electronic structure. Merz’s lab saw the potential of SQD after IBM scientists presented the algorithm a few years ago. The resulting workflow has already demonstrated competitive accuracy compared to the most demanding classical approaches, and researchers believe it can scale to even larger molecules. Merz hopes to build databases of simulated molecular behavior that can be used with machine learning to identify molecules that might behave in certain ways, and then synthesize them.
It addresses one of the fundamental challenges of electronic structure calculations: the number of possible configurations of a molecule’s electrons grows combinatorially with the molecule’s size.
Hybrid Quantum-Classical Workflows Scale Towards Molecular Databases
The ability to accurately simulate molecular behavior promises to accelerate discovery across diverse fields, and a recent collaboration between Cleveland Clinic and IBM has demonstrated a significant step toward that goal. This achievement addresses a longstanding challenge in computational chemistry; as molecular size increases, the computational demands of accurate electronic structure calculations on classical computers escalate dramatically. Simpler clusters can be efficiently modeled using classical methods. This isn’t simply about improving existing simulations, but rather establishing a foundation for quantum-centric supercomputing (QCSC). Merz envisions a future where these workflows support the creation of databases of simulated molecular behavior, enabling scientists to utilize machine learning to identify molecules that might behave in desired ways.
Proving that this approach works for Trp-cage is a step toward larger molecules.
Mario Motta, co-author of the paper