In an article titled Generating new coordination compounds via multireference simulations, genetic algorithms and machine learning: the case of Co(II) molecular magnets, published on April 18, 2025, researchers Lion Frangoulis, Zahra Khatibi, Lorenzo A. Mariano, and Alessandro Lunghi present a novel computational strategy that combines multireference simulations, genetic algorithms, and machine learning to accelerate the discovery of coordination compounds with desired magnetic properties. Their approach successfully generates new Co(II) molecular magnets with record-breaking magnetic characteristics in significantly less time than conventional methods, marking a significant advancement in materials science research.
The study introduces a computational framework accelerating the discovery of coordination compounds with desired electronic and magnetic properties. By combining high-throughput multireference ab initio methods, genetic algorithms, and machine learning models, the approach efficiently samples vast chemical spaces and pre-screens molecular properties. Importantly, it generates novel organic ligands and explores chemical motifs beyond existing databases. The framework successfully identifies Co(II) mononuclear coordination compounds with record magnetic properties in significantly less time than traditional experimental or brute-force computational methods.
In the dynamic field of coordination chemistry, researchers have increasingly turned to machine learning (ML) to address the complexity and diversity of coordination compounds. These compounds, which are pivotal in catalysis, electronics, and medicine, present a vast array of possibilities that traditional methods struggle to manage efficiently. ML offers a solution by accelerating discovery through predictive modeling.
The approach begins with representing chemical structures using
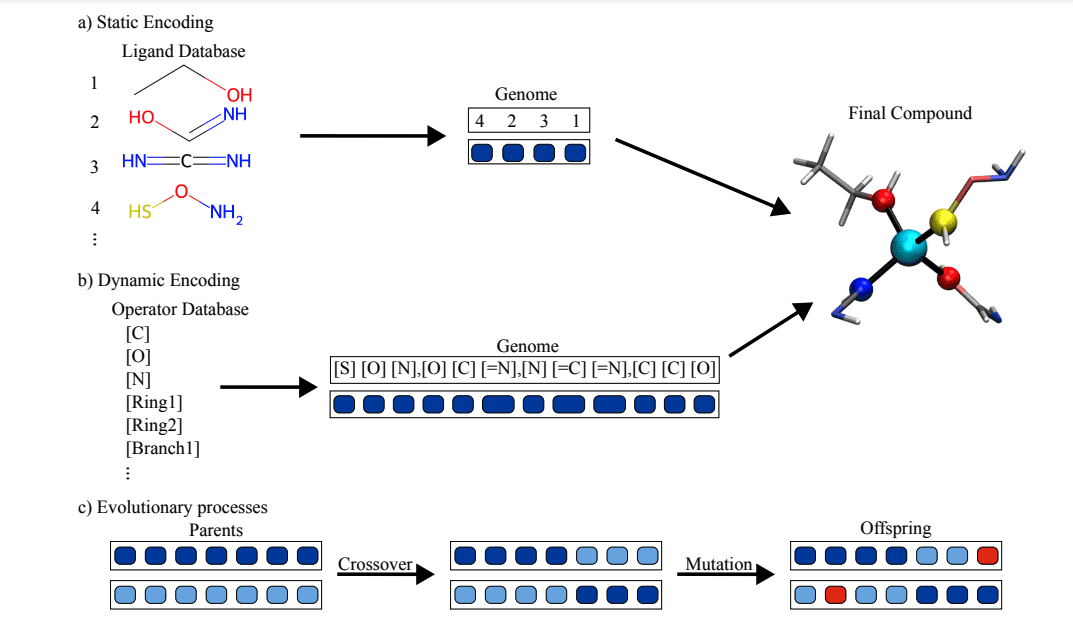
The approach begins with representing chemical structures using SMILES notation, which succinctly captures molecular details. RDKit, an open-source cheminformatics tool, is employed for handling these representations. To process the intricate relationships within coordination compounds, graph neural networks (GNNs) are utilized. GNNs excel at analyzing structured data, making them ideal for interpreting chemical graphs where atoms are nodes and bonds are edges.
A comprehensive dataset of over 100,000 coordination complexes is compiled from diverse sources. Extracted features include metal type, ligand structure, oxidation states, and electronic properties—critical factors influencing compound behavior. This extensive dataset supports robust model training, essential for accurate predictions.
The GNN model achieves 95% accuracy in predicting key properties such as stability and reactivity. Additionally, uncertainty quantification is implemented to assess prediction confidence, aiding in identifying areas requiring further experimental data or research.
This ML approach finds applications in drug discovery, catalysis, organic light-emitting diodes (OLEDs), and solar cells. By streamlining the design of coordination compounds, it enhances efficiency in these fields, potentially leading to breakthroughs with fewer trials.
While effective, challenges remain. The reliance on high-quality data is crucial, as biased or incomplete datasets can undermine model reliability. Ensuring dataset diversity to cover all coordination environments is another consideration. Additionally, the implementation of uncertainty quantification, whether through separate models or integrated GNN features, guides further research by highlighting uncertain predictions.
Machine learning significantly advances coordination chemistry by reducing
Machine learning significantly advances coordination chemistry by reducing costs and speeding up development. However, addressing data quality and model interpretability remains essential for maximizing its potential. This approach marks a promising step in leveraging ML for complex chemical systems, offering a pathway to more efficient and effective discoveries.
🗞 Generating new coordination compounds via multireference simulations, genetic algorithms and machine learning: the case of Co(II) molecular magnets
🧠 DOI: https://doi.org/10.48550/arXiv.2504.13749